Dr. Dominic Grün (CIBSS-AI), Max Planck Institute of Immunobiology and Epigenetics Freiburg
The liver is the major metabolic organ of the body and exerts central physiological functions. Not surprisingly, liver malignancies such as fatty liver disease, Hepatitis, and liver cancer represent a major burden for the society. The liver exhibits an extraordinary regenerative capacity in response to liver damage, which is implicated in disease pathogenesis. However, the cellular and molecular processes of liver regeneration are incompletely understood. With the advent of single-cell RNA-sequencing technologies it became possible to investigate the liver at single-cell resolution, leading to major novel insights into cell type heterogeneity and spatial zonation of the liver. However, detailed knowledge on the effect of physical and signalling interactions of liver cell types on metabolic function and cell fate control in the liver during health and disease is still lacking.
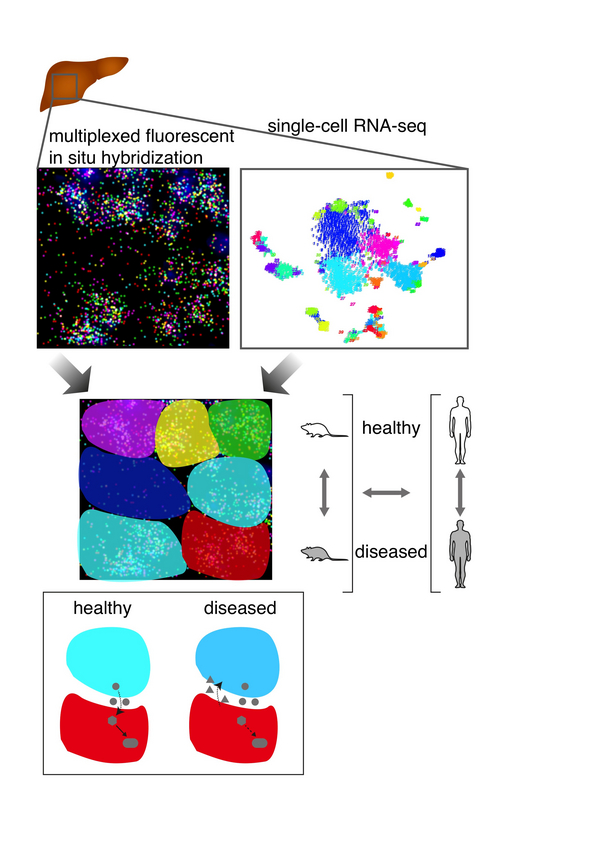
In this project we will follow-up on our recent work on the characterization of human liver cell types with a specific focus on liver stem cells. To explore similarities and differences between human liver and the mouse model we will first create a comprehensive mouse liver cell type atlas and quantify evolutionary conservation of cell type-specific programs between human and mouse. In a second step we will investigate spatial co-localization of liver cell types and identify pathways of cellular crosstalk during homeostasis and regeneration in the mouse model. To perform this spatial high-resolution analysis we will combine the power of single-cell RNA-seq for the identification of cell types with the simultaneous quantification of large numbers of genes at single mRNA molecule resolution in tissue sections with the help of advanced single-molecule FISH methods. Inference of signalling pathways for cell-cell communication and computational modelling of differentiation trajectories and regulatory networks of interacting cell types will then allow us to explore the role of cell-cell communication in controlling cell states in the liver.